|
However, the efflux function of cells can exclude GNRs extracellularly within 24 h, resulting in reduced cell mitochondrial metabolic toxicity and allowing metabolic disorders to recover to normal function.Īdipocytes or fat cells play a vital role in the storage and release of energy in pigs, and many circular RNAs (circRNAs) have emerged as important regulators in various tissues and cell types in pigs. The mitochondrial dysfunction disrupted alanine, aspartate, glutamate, arginine, and proline metabolism, with amino acid synthesis generally downregulated. The cytotoxicity and bioTEM results illustrated that the mitochondria toxicity, as the main cytotoxicity of GNRs, caused declining cytoprotective ability. 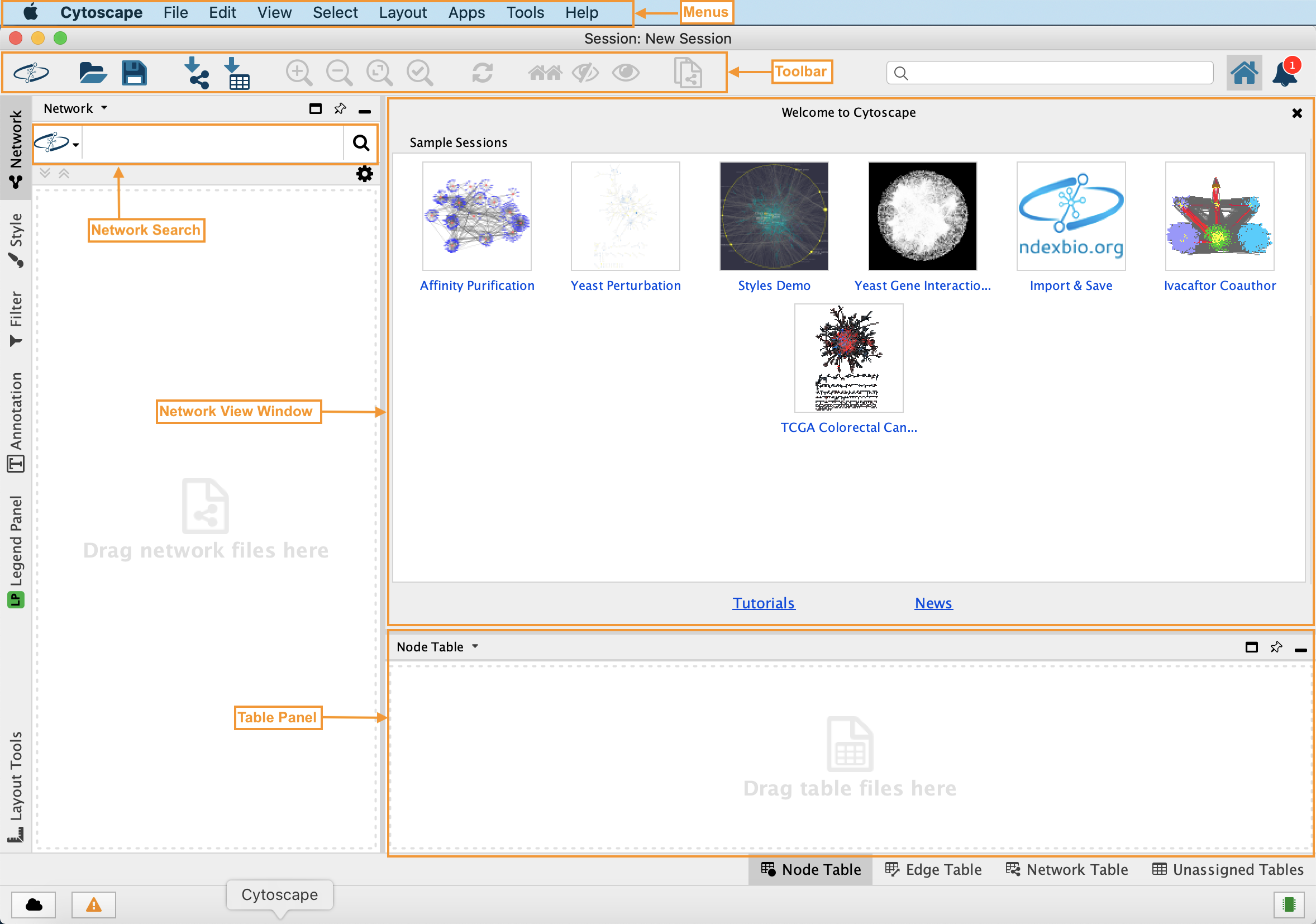
High concentrations of GNRs caused more significant toxicity. The shape of GNRs affected cytotoxicity, and short GNR (GNR−S) triggered disorder of cell metabolism. Key metabolic pathways affected by GNRs were identified by endo-metabolic profiling of cells mixed with GNRs of varying shape while varying the dose and time of exposure. Using optimized parameters for chromatography and mass spectrometry, the metabolites detected by GC-MS were processed with MS DIAL and identified with Fiehnlib. In this study, the human hepatoma cell line (Hep G2) was used in a metabolomics approach to study the effects of shape, time, and dose of gold nanorods (GNRs). 2011 February 1 27(3): 431–432.Published online 2010 December 12.Despite the increasing application of gold nanoparticles, there has been little assessment of biological system toxicity to evaluate their potential impact on human health. This experiment uses : Cytoscape, Smoot, Keiichiro Ono, Johannes Ruscheinski, Peng-Liang Wang, Trey Ideker, Cytoscape 2.8: new features for data integration and network visualization Bioinformatics. The chart can be exported and can be saved as jpeg file. Shows the distribution of the degree with number of nodes, there are different options to enlarge chart, to export the chart, and to fit a line. The simple network parameters, analyze the network density, network radius, clustering coefficient, number of nodes and self loops, etc.Ĭomplex network parameter shows the distribution of node, shortest path length, topological coefficient distribution etc.įigure 7: screenshot of node degree distribution Select according to the network one have downloaded.įigure 5: screebshot to interpret the networkĪfter one has chosen the network interpretation the network is analyzed with simple and complex parameters.įigure 6: Scrrenshot of the results with simple parameters When one chooses to analyze network, the GUI appears asking the user to interpret network as undirected or directed network. Step 6: Go to plug in menu, choose the option "Network Analysis", followed by choose "Analyze network". Step 5: Once the network is loaded, change it into preferred layout.įigure 3: Screen shot to manage with layouts Step 4: Import the network into the cytoscape, by the standard menu "File" option as shown below.įigure 2: Screenshot to import network files One can download the network from external databases or can also import the network directly from the web services. Step 3: Go to the required pathway database exaple IntAct, download the network in a suitable format like sif, sbml, xml, etc. Go to plug-in options in the standard menu bar, check whether the network analyzer is installed or not, if not go to manage plug in and search for Network analyzer and install.įigure 1: Screenshot of Cytoscape and to manage plugin menu 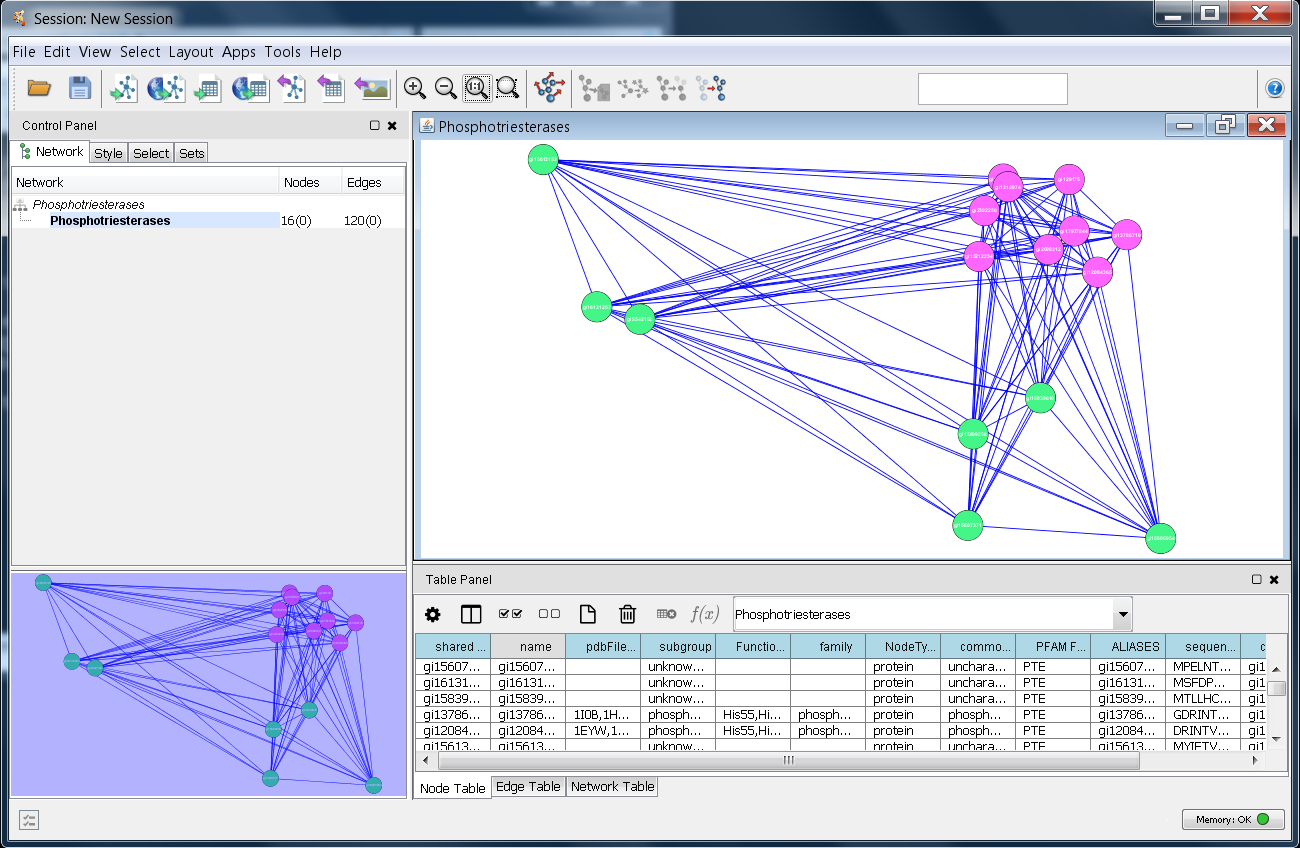
Step 2: Launch cytoscape, make sure that you have internet in your computer. Please go to simulator tab for detailed information about instalation. 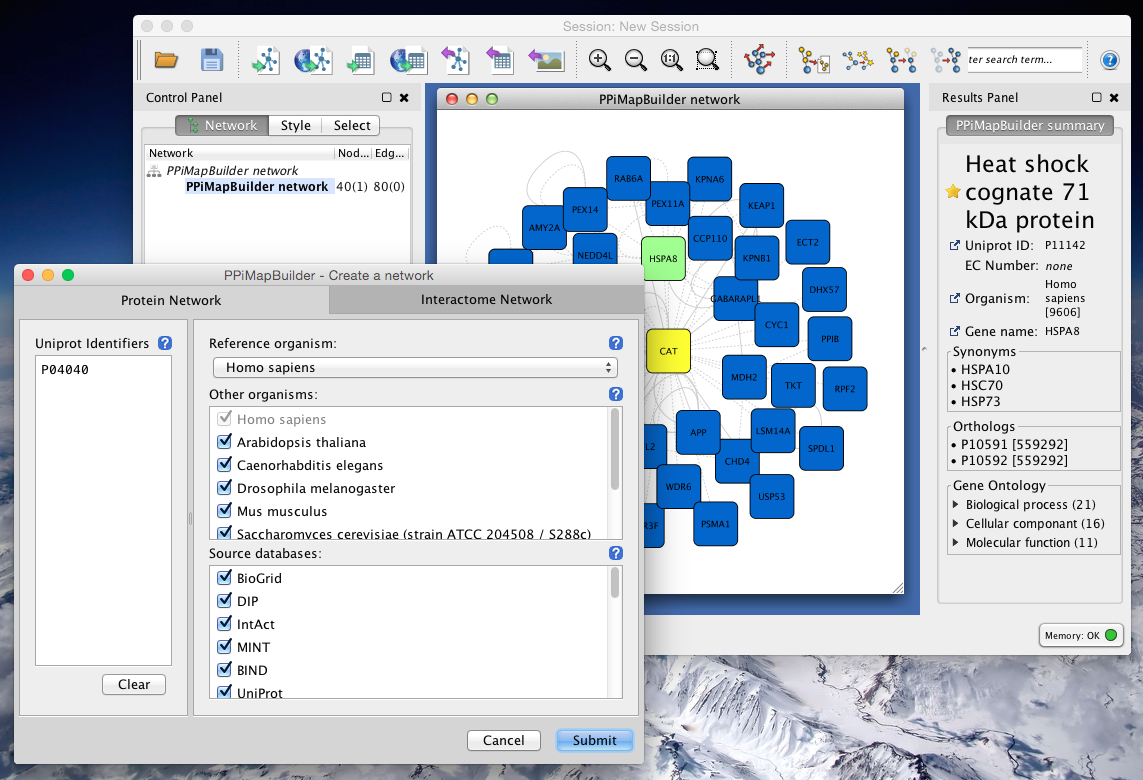
Step1: Download and install cytoscape in your computer. Follow the steps given to analyse the network and feature detection
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
